Search Results for author: Seyed-Ahmad Ahmadi
Found 23 papers, 9 papers with code
MedShapeNet -- A Large-Scale Dataset of 3D Medical Shapes for Computer Vision
1 code implementation • 30 Aug 2023 • Jianning Li, Zongwei Zhou, Jiancheng Yang, Antonio Pepe, Christina Gsaxner, Gijs Luijten, Chongyu Qu, Tiezheng Zhang, Xiaoxi Chen, Wenxuan Li, Marek Wodzinski, Paul Friedrich, Kangxian Xie, Yuan Jin, Narmada Ambigapathy, Enrico Nasca, Naida Solak, Gian Marco Melito, Viet Duc Vu, Afaque R. Memon, Christopher Schlachta, Sandrine de Ribaupierre, Rajnikant Patel, Roy Eagleson, Xiaojun Chen, Heinrich Mächler, Jan Stefan Kirschke, Ezequiel de la Rosa, Patrick Ferdinand Christ, Hongwei Bran Li, David G. Ellis, Michele R. Aizenberg, Sergios Gatidis, Thomas Küstner, Nadya Shusharina, Nicholas Heller, Vincent Andrearczyk, Adrien Depeursinge, Mathieu Hatt, Anjany Sekuboyina, Maximilian Löffler, Hans Liebl, Reuben Dorent, Tom Vercauteren, Jonathan Shapey, Aaron Kujawa, Stefan Cornelissen, Patrick Langenhuizen, Achraf Ben-Hamadou, Ahmed Rekik, Sergi Pujades, Edmond Boyer, Federico Bolelli, Costantino Grana, Luca Lumetti, Hamidreza Salehi, Jun Ma, Yao Zhang, Ramtin Gharleghi, Susann Beier, Arcot Sowmya, Eduardo A. Garza-Villarreal, Thania Balducci, Diego Angeles-Valdez, Roberto Souza, Leticia Rittner, Richard Frayne, Yuanfeng Ji, Vincenzo Ferrari, Soumick Chatterjee, Florian Dubost, Stefanie Schreiber, Hendrik Mattern, Oliver Speck, Daniel Haehn, Christoph John, Andreas Nürnberger, João Pedrosa, Carlos Ferreira, Guilherme Aresta, António Cunha, Aurélio Campilho, Yannick Suter, Jose Garcia, Alain Lalande, Vicky Vandenbossche, Aline Van Oevelen, Kate Duquesne, Hamza Mekhzoum, Jef Vandemeulebroucke, Emmanuel Audenaert, Claudia Krebs, Timo Van Leeuwen, Evie Vereecke, Hauke Heidemeyer, Rainer Röhrig, Frank Hölzle, Vahid Badeli, Kathrin Krieger, Matthias Gunzer, Jianxu Chen, Timo van Meegdenburg, Amin Dada, Miriam Balzer, Jana Fragemann, Frederic Jonske, Moritz Rempe, Stanislav Malorodov, Fin H. Bahnsen, Constantin Seibold, Alexander Jaus, Zdravko Marinov, Paul F. Jaeger, Rainer Stiefelhagen, Ana Sofia Santos, Mariana Lindo, André Ferreira, Victor Alves, Michael Kamp, Amr Abourayya, Felix Nensa, Fabian Hörst, Alexander Brehmer, Lukas Heine, Yannik Hanusrichter, Martin Weßling, Marcel Dudda, Lars E. Podleska, Matthias A. Fink, Julius Keyl, Konstantinos Tserpes, Moon-Sung Kim, Shireen Elhabian, Hans Lamecker, Dženan Zukić, Beatriz Paniagua, Christian Wachinger, Martin Urschler, Luc Duong, Jakob Wasserthal, Peter F. Hoyer, Oliver Basu, Thomas Maal, Max J. H. Witjes, Gregor Schiele, Ti-chiun Chang, Seyed-Ahmad Ahmadi, Ping Luo, Bjoern Menze, Mauricio Reyes, Thomas M. Deserno, Christos Davatzikos, Behrus Puladi, Pascal Fua, Alan L. Yuille, Jens Kleesiek, Jan Egger
For the medical domain, we present a large collection of anatomical shapes (e. g., bones, organs, vessels) and 3D models of surgical instrument, called MedShapeNet, created to facilitate the translation of data-driven vision algorithms to medical applications and to adapt SOTA vision algorithms to medical problems.
DIAMANT: Dual Image-Attention Map Encoders For Medical Image Segmentation
no code implementations • 28 Apr 2023 • Yousef Yeganeh, Azade Farshad, Peter Weinberger, Seyed-Ahmad Ahmadi, Ehsan Adeli, Nassir Navab
Although purely transformer-based architectures showed promising performance in many computer vision tasks, many hybrid models consisting of CNN and transformer blocks are introduced to fit more specialized tasks.
Open-Source Skull Reconstruction with MONAI
1 code implementation • 25 Nov 2022 • Jianning Li, André Ferreira, Behrus Puladi, Victor Alves, Michael Kamp, Moon-Sung Kim, Felix Nensa, Jens Kleesiek, Seyed-Ahmad Ahmadi, Jan Egger
The primary goal of this paper lies in the investigation of open-sourcing codes and pre-trained deep learning models under the MONAI framework.
Training β-VAE by Aggregating a Learned Gaussian Posterior with a Decoupled Decoder
1 code implementation • 29 Sep 2022 • Jianning Li, Jana Fragemann, Seyed-Ahmad Ahmadi, Jens Kleesiek, Jan Egger
The reconstruction loss and the Kullback-Leibler divergence (KLD) loss in a variational autoencoder (VAE) often play antagonistic roles, and tuning the weight of the KLD loss in $\beta$-VAE to achieve a balance between the two losses is a tricky and dataset-specific task.
Graph-in-Graph (GiG): Learning interpretable latent graphs in non-Euclidean domain for biological and healthcare applications
no code implementations • 1 Apr 2022 • Kamilia Mullakaeva, Luca Cosmo, Anees Kazi, Seyed-Ahmad Ahmadi, Nassir Navab, Michael M. Bronstein
In this work, we propose Graph-in-Graph (GiG), a neural network architecture for protein classification and brain imaging applications that exploits the graph representation of the input data samples and their latent relation.
Simultaneous imputation and disease classification in incomplete medical datasets using Multigraph Geometric Matrix Completion (MGMC)
1 code implementation • 14 May 2020 • Gerome Vivar, Anees Kazi, Hendrik Burwinkel, Andreas Zwergal, Nassir Navab, Seyed-Ahmad Ahmadi
As a solution, we propose an end-to-end learning of imputation and disease prediction of incomplete medical datasets via Multigraph Geometric Matrix Completion (MGMC).
Domain-specific loss design for unsupervised physical training: A new approach to modeling medical ML solutions
no code implementations • 9 May 2020 • Hendrik Burwinkel, Holger Matz, Stefan Saur, Christoph Hauger, Ayse Mine Evren, Nino Hirnschall, Oliver Findl, Nassir Navab, Seyed-Ahmad Ahmadi
The cataract, a developing opacity of the human eye lens, constitutes the world's most frequent cause for blindness.
Decision Support for Intoxication Prediction Using Graph Convolutional Networks
no code implementations • 2 May 2020 • Hendrik Burwinkel, Matthias Keicher, David Bani-Harouni, Tobias Zellner, Florian Eyer, Nassir Navab, Seyed-Ahmad Ahmadi
Due to the time-sensitive nature of these cases, doctors are required to propose a correct diagnosis and intervention within a minimal time frame.
Peri-Diagnostic Decision Support Through Cost-Efficient Feature Acquisition at Test-Time
no code implementations • 31 Mar 2020 • Gerome Vivar, Kamilia Mullakaeva, Andreas Zwergal, Nassir Navab, Seyed-Ahmad Ahmadi
Computer-aided diagnosis (CADx) algorithms in medicine provide patient-specific decision support for physicians.
Latent-Graph Learning for Disease Prediction
no code implementations • 27 Mar 2020 • Luca Cosmo, Anees Kazi, Seyed-Ahmad Ahmadi, Nassir Navab, Michael Bronstein
Recently, Graph Convolutional Networks (GCNs) have proven to be a powerful machine learning tool for Computer-Aided Diagnosis (CADx) and disease prediction.
Differentiable Graph Module (DGM) for Graph Convolutional Networks
1 code implementation • 11 Feb 2020 • Anees Kazi, Luca Cosmo, Seyed-Ahmad Ahmadi, Nassir Navab, Michael Bronstein
We provide an extensive evaluation of applications from the domains of healthcare (disease prediction), brain imaging (age prediction), computer graphics (3D point cloud segmentation), and computer vision (zero-shot learning).
Multi-modal Graph Fusion for Inductive Disease Classification in Incomplete Datasets
no code implementations • 8 May 2019 • Gerome Vivar, Hendrik Burwinkel, Anees Kazi, Andreas Zwergal, Nassir Navab, Seyed-Ahmad Ahmadi
Recently, several works proposed geometric deep learning approaches to solve disease classification, by modeling patients as nodes in a graph, along with graph signal processing of multi-modal features.
Adaptive Image-Feature Learning for Disease Classification Using Inductive Graph Networks
no code implementations • 8 May 2019 • Hendrik Burwinkel, Anees Kazi, Gerome Vivar, Shadi Albarqouni, Guillaume Zahnd, Nassir Navab, Seyed-Ahmad Ahmadi
We propose a new network architecture that exploits an inductive end-to-end learning approach for disease classification, where filters from both the CNN and the graph are trained jointly.
InceptionGCN: Receptive Field Aware Graph Convolutional Network for Disease Prediction
no code implementations • 11 Mar 2019 • Anees Kazi, Shayan shekarforoush, S. Arvind krishna, Hendrik Burwinkel, Gerome Vivar, Karsten Kortuem, Seyed-Ahmad Ahmadi, Shadi Albarqouni, Nassir Navab
Geometric deep learning provides a principled and versatile manner for the integration of imaging and non-imaging modalities in the medical domain.
Stabilizing Inputs to Approximated Nonlinear Functions for Inference with Homomorphic Encryption in Deep Neural Networks
no code implementations • 5 Feb 2019 • Moustafa AboulAtta, Matthias Ossadnik, Seyed-Ahmad Ahmadi
Leveled Homomorphic Encryption (LHE) offers a potential solution that could allow sectors with sensitive data to utilize the cloud and securely deploy their models for remote inference with Deep Neural Networks (DNN).
Multi-modal Disease Classification in Incomplete Datasets Using Geometric Matrix Completion
no code implementations • 30 Mar 2018 • Gerome Vivar, Andreas Zwergal, Nassir Navab, Seyed-Ahmad Ahmadi
In this work, we follow up on the idea of modeling multi-modal disease classification as a matrix completion problem, with simultaneous classification and non-linear imputation of features.
TOMAAT: volumetric medical image analysis as a cloud service
no code implementations • 19 Mar 2018 • Fausto Milletari, Johann Frei, Seyed-Ahmad Ahmadi
Deep learning has been recently applied to a multitude of computer vision and medical image analysis problems.
Classification of sparsely labeled spatio-temporal data through semi-supervised adversarial learning
no code implementations • 26 Jan 2018 • Atanas Mirchev, Seyed-Ahmad Ahmadi
In recent years, Generative Adversarial Networks (GAN) have emerged as a powerful method for learning the mapping from noisy latent spaces to realistic data samples in high-dimensional space.
SurvivalNet: Predicting patient survival from diffusion weighted magnetic resonance images using cascaded fully convolutional and 3D convolutional neural networks
1 code implementation • 20 Feb 2017 • Patrick Ferdinand Christ, Florian Ettlinger, Georgios Kaissis, Sebastian Schlecht, Freba Ahmaddy, Felix Grün, Alexander Valentinitsch, Seyed-Ahmad Ahmadi, Rickmer Braren, Bjoern Menze
A 3D neural network (SurvivalNet) then predicts the HCC lesions' malignancy from the HCC tumor segmentation.
Automatic Liver and Tumor Segmentation of CT and MRI Volumes using Cascaded Fully Convolutional Neural Networks
4 code implementations • 20 Feb 2017 • Patrick Ferdinand Christ, Florian Ettlinger, Felix Grün, Mohamed Ezzeldin A. Elshaera, Jana Lipkova, Sebastian Schlecht, Freba Ahmaddy, Sunil Tatavarty, Marc Bickel, Patrick Bilic, Markus Rempfler, Felix Hofmann, Melvin D Anastasi, Seyed-Ahmad Ahmadi, Georgios Kaissis, Julian Holch, Wieland Sommer, Rickmer Braren, Volker Heinemann, Bjoern Menze
In the first step, we train a FCN to segment the liver as ROI input for a second FCN.
 Automatic Liver And Tumor Segmentation
Automatic Liver And Tumor Segmentation
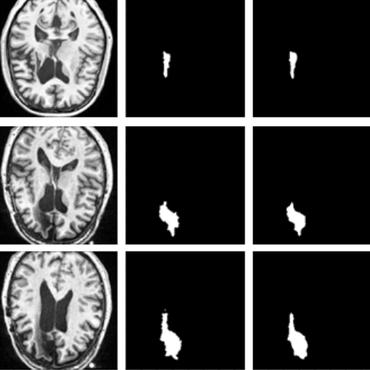 Lesion Segmentation
+2
Lesion Segmentation
+2
Automatic Liver and Lesion Segmentation in CT Using Cascaded Fully Convolutional Neural Networks and 3D Conditional Random Fields
3 code implementations • 7 Oct 2016 • Patrick Ferdinand Christ, Mohamed Ezzeldin A. Elshaer, Florian Ettlinger, Sunil Tatavarty, Marc Bickel, Patrick Bilic, Markus Rempfler, Marco Armbruster, Felix Hofmann, Melvin D'Anastasi, Wieland H. Sommer, Seyed-Ahmad Ahmadi, Bjoern H. Menze
Automatic segmentation of the liver and its lesion is an important step towards deriving quantitative biomarkers for accurate clinical diagnosis and computer-aided decision support systems.
V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation
27 code implementations • 15 Jun 2016 • Fausto Milletari, Nassir Navab, Seyed-Ahmad Ahmadi
Convolutional Neural Networks (CNNs) have been recently employed to solve problems from both the computer vision and medical image analysis fields.
Hough-CNN: Deep Learning for Segmentation of Deep Brain Regions in MRI and Ultrasound
no code implementations • 26 Jan 2016 • Fausto Milletari, Seyed-Ahmad Ahmadi, Christine Kroll, Annika Plate, Verena Rozanski, Juliana Maiostre, Johannes Levin, Olaf Dietrich, Birgit Ertl-Wagner, Kai Bötzel, Nassir Navab
In this work we propose a novel approach to perform segmentation by leveraging the abstraction capabilities of convolutional neural networks (CNNs).





