BioT5+: Towards Generalized Biological Understanding with IUPAC Integration and Multi-task Tuning
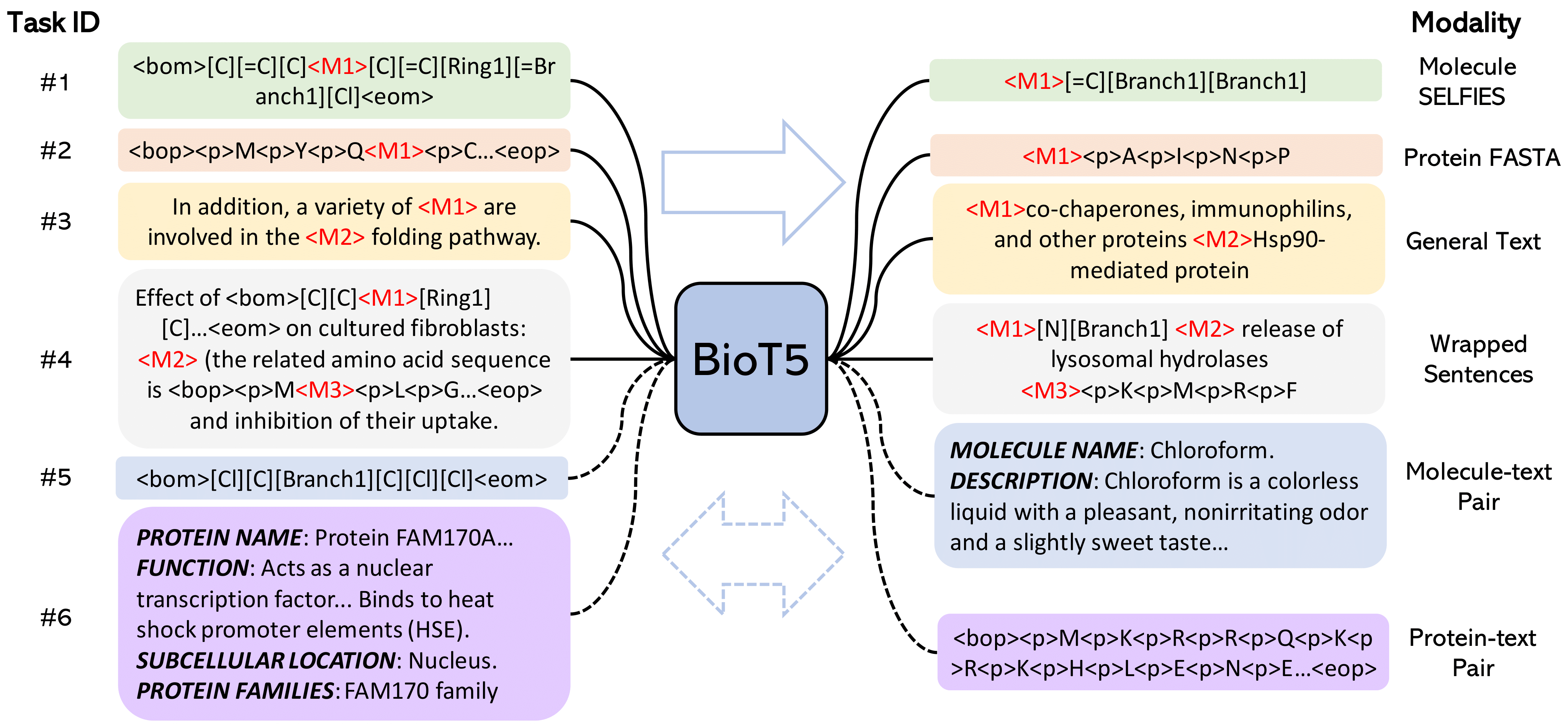
Recent research trends in computational biology have increasingly focused on integrating text and bio-entity modeling, especially in the context of molecules and proteins. However, previous efforts like BioT5 faced challenges in generalizing across diverse tasks and lacked a nuanced understanding of molecular structures, particularly in their textual representations (e.g., IUPAC). This paper introduces BioT5+, an extension of the BioT5 framework, tailored to enhance biological research and drug discovery. BioT5+ incorporates several novel features: integration of IUPAC names for molecular understanding, inclusion of extensive bio-text and molecule data from sources like bioRxiv and PubChem, the multi-task instruction tuning for generality across tasks, and a novel numerical tokenization technique for improved processing of numerical data. These enhancements allow BioT5+ to bridge the gap between molecular representations and their textual descriptions, providing a more holistic understanding of biological entities, and largely improving the grounded reasoning of bio-text and bio-sequences. The model is pre-trained and fine-tuned with a large number of experiments, including \emph{3 types of problems (classification, regression, generation), 15 kinds of tasks, and 21 total benchmark datasets}, demonstrating the remarkable performance and state-of-the-art results in most cases. BioT5+ stands out for its ability to capture intricate relationships in biological data, thereby contributing significantly to bioinformatics and computational biology. Our code is available at \url{https://github.com/QizhiPei/BioT5}.
PDF AbstractCode
Datasets
| Task | Dataset | Model | Metric Name | Metric Value | Global Rank | Benchmark |
|---|---|---|---|---|---|---|
| Molecule Captioning | ChEBI-20 | BioT5+ | BLEU-2 | 66.6 | # 1 | |
| BLEU-4 | 59.1 | # 1 | ||||
| ROUGE-1 | 71.0 | # 1 | ||||
| ROUGE-2 | 58.4 | # 1 | ||||
| ROUGE-L | 65.0 | # 1 | ||||
| METEOR | 68.1 | # 1 | ||||
| Text-based de novo Molecule Generation | ChEBI-20 | BioT5+ | Text2Mol | 57.9 | # 6 | |
| BLEU | 87.2 | # 1 | ||||
| Exact Match | 52.2 | # 1 | ||||
| Levenshtein | 12.776 | # 16 | ||||
| MACCS FTS | 90.7 | # 1 | ||||
| RDK FTS | 83.5 | # 1 | ||||
| Morgan FTS | 77.9 | # 1 | ||||
| Frechet ChemNet Distance (FCD) | 0.353 | # 5 | ||||
| Validity | 100 | # 1 | ||||
| Parameter Count | 252000000 | # 13 | ||||
| Retrosynthesis | Mol-Instruction | BioT5+ | Exact | 0.642 | # 2 | |
| Validity | 1 | # 1 | ||||
| Morgan FTS | 0.866 | # 2 | ||||
| Reagent Prediction | Mol-Instruction | BioT5+ | Exact | 0.257 | # 2 | |
| Validity | 1 | # 1 | ||||
| Morgan FTS | 0.512 | # 2 | ||||
| Forward reaction prediction | Mol-Instruction | BioT5+ | Exact | 0.864 | # 2 | |
| Validity | 1 | # 1 | ||||
| Morgan FTS | 0.935 | # 1 |








 MoleculeNet
MoleculeNet
 QM9
QM9
 ChEBI-20
ChEBI-20
