Search Results for author: Bruce Fischl
Found 27 papers, 11 papers with code
Boosting Skull-Stripping Performance for Pediatric Brain Images
no code implementations • 26 Feb 2024 • William Kelley, Nathan Ngo, Adrian V. Dalca, Bruce Fischl, Lilla Zöllei, Malte Hoffmann
With the emergence of multi-institutional pediatric data acquisition efforts to broaden the understanding of perinatal brain development, it is essential to develop robust and well-tested tools ready for the relevant data processing.
Fully Convolutional Slice-to-Volume Reconstruction for Single-Stack MRI
1 code implementation • 5 Dec 2023 • Sean I. Young, Yaël Balbastre, Bruce Fischl, Polina Golland, Juan Eugenio Iglesias
Here, we propose a SVR method that overcomes the shortcomings of previous work and produces state-of-the-art reconstructions in the presence of extreme inter-slice motion.
JOSA: Joint surface-based registration and atlas construction of brain geometry and function
no code implementations • 22 Oct 2023 • Jian Li, Greta Tuckute, Evelina Fedorenko, Brian L. Edlow, Adrian V. Dalca, Bruce Fischl
By recognizing the mismatch between geometry and function, JOSA provides new insights into the future development of registration methods using joint analysis of the brain structure and function.
Multi-Head Graph Convolutional Network for Structural Connectome Classification
no code implementations • 2 May 2023 • Anees Kazi, Jocelyn Mora, Bruce Fischl, Adrian V. Dalca, Iman Aganj
To test the ability of our model to extract complementary and representative features from brain connectivity data, we chose the task of sex classification.
Joint cortical registration of geometry and function using semi-supervised learning
no code implementations • 2 Mar 2023 • Jian Li, Greta Tuckute, Evelina Fedorenko, Brian L. Edlow, Bruce Fischl, Adrian V. Dalca
Brain surface-based image registration, an important component of brain image analysis, establishes spatial correspondence between cortical surfaces.
Anatomy-aware and acquisition-agnostic joint registration with SynthMorph
no code implementations • 26 Jan 2023 • Malte Hoffmann, Andrew Hoopes, Douglas N. Greve, Bruce Fischl, Adrian V. Dalca
Most affine methods are agnostic to anatomy, meaning the registration will be inaccurate if algorithms consider all structures in the image.
Data Consistent Deep Rigid MRI Motion Correction
1 code implementation • 25 Jan 2023 • Nalini M. Singh, Neel Dey, Malte Hoffmann, Bruce Fischl, Elfar Adalsteinsson, Robert Frost, Adrian V. Dalca, Polina Golland
Motion artifacts are a pervasive problem in MRI, leading to misdiagnosis or mischaracterization in population-level imaging studies.
SuperWarp: Supervised Learning and Warping on U-Net for Invariant Subvoxel-Precise Registration
no code implementations • 15 May 2022 • Sean I. Young, Yaël Balbastre, Adrian V. Dalca, William M. Wells, Juan Eugenio Iglesias, Bruce Fischl
In recent years, learning-based image registration methods have gradually moved away from direct supervision with target warps to instead use self-supervision, with excellent results in several registration benchmarks.
Learning the Effect of Registration Hyperparameters with HyperMorph
no code implementations • 30 Mar 2022 • Andrew Hoopes, Malte Hoffmann, Douglas N. Greve, Bruce Fischl, John Guttag, Adrian V. Dalca
We design a meta network, or hypernetwork, that predicts the parameters of a registration network for input hyperparameters, thereby comprising a single model that generates the optimal deformation field corresponding to given hyperparameter values.
SynthStrip: Skull-Stripping for Any Brain Image
no code implementations • 18 Mar 2022 • Andrew Hoopes, Jocelyn S. Mora, Adrian V. Dalca, Bruce Fischl, Malte Hoffmann
The removal of non-brain signal from magnetic resonance imaging (MRI) data, known as skull-stripping, is an integral component of many neuroimage analysis streams.
Supervision by Denoising for Medical Image Segmentation
no code implementations • 7 Feb 2022 • Sean I. Young, Adrian V. Dalca, Enzo Ferrante, Polina Golland, Christopher A. Metzler, Bruce Fischl, Juan Eugenio Iglesias
SUD unifies stochastic averaging and spatial denoising techniques under a spatio-temporal denoising framework and alternates denoising and model weight update steps in an optimization framework for semi-supervision.
SynthSeg: Segmentation of brain MRI scans of any contrast and resolution without retraining
2 code implementations • 20 Jul 2021 • Benjamin Billot, Douglas N. Greve, Oula Puonti, Axel Thielscher, Koen van Leemput, Bruce Fischl, Adrian V. Dalca, Juan Eugenio Iglesias
Here we introduce SynthSeg, the first segmentation CNN robust against changes in contrast and resolution.
Unsupervised learning of MRI tissue properties using MRI physics models
no code implementations • 6 Jul 2021 • Divya Varadarajan, Katherine L. Bouman, Andre van der Kouwe, Bruce Fischl, Adrian V. Dalca
In this work we propose an unsupervised deep-learning strategy that employs MRI physics to estimate all three tissue properties from a single multiecho MRI scan session, and generalizes across varying acquisition parameters.
Robust joint registration of multiple stains and MRI for multimodal 3D histology reconstruction: Application to the Allen human brain atlas
1 code implementation • 30 Apr 2021 • Adrià Casamitjana, Marco Lorenzi, Sebastiano Ferraris, Loc Peter, Marc Modat, Allison Stevens, Bruce Fischl, Tom Vercauteren, Juan Eugenio Iglesias
The model relies on a spanning tree of latent transforms connecting all the sections and slices of the reference volume, and assumes that the registration between any pair of images can be see as a noisy version of the composition of (possibly inverted) latent transforms connecting the two images.
HyperMorph: Amortized Hyperparameter Learning for Image Registration
1 code implementation • 4 Jan 2021 • Andrew Hoopes, Malte Hoffmann, Bruce Fischl, John Guttag, Adrian V. Dalca
We present HyperMorph, a learning-based strategy for deformable image registration that removes the need to tune important registration hyperparameters during training.
Joint super-resolution and synthesis of 1 mm isotropic MP-RAGE volumes from clinical MRI exams with scans of different orientation, resolution and contrast
1 code implementation • 24 Dec 2020 • Juan Eugenio Iglesias, Benjamin Billot, Yael Balbastre, Azadeh Tabari, John Conklin, Daniel C. Alexander, Polina Golland, Brian L. Edlow, Bruce Fischl
Most existing algorithms for automatic 3D morphometry of human brain MRI scans are designed for data with near-isotropic voxels at approximately 1 mm resolution, and frequently have contrast constraints as well - typically requiring T1 scans (e. g., MP-RAGE).
3D Reconstruction and Segmentation of Dissection Photographs for MRI-free Neuropathology
1 code implementation • 11 Sep 2020 • Henry Tregidgo, Adria Casamitjana, Caitlin Latimer, Mitchell Kilgore, Eleanor Robinson, Emily Blackburn, Koen van Leemput, Bruce Fischl, Adrian Dalca, Christine Mac Donald, Dirk Keene, Juan Eugenio Iglesias
NTNC traditionally requires a volumetric MRI scan, acquired either ex vivo or a short time prior to death.
SynthMorph: learning contrast-invariant registration without acquired images
no code implementations • 21 Apr 2020 • Malte Hoffmann, Benjamin Billot, Douglas N. Greve, Juan Eugenio Iglesias, Bruce Fischl, Adrian V. Dalca
This approach results in powerful networks that accurately generalize to a broad array of MRI contrasts.
Cortical surface registration using unsupervised learning
1 code implementation • 9 Apr 2020 • Jieyu Cheng, Adrian V. Dalca, Bruce Fischl, Lilla Zollei
The experiments show that the proposed SphereMorph is capable of modeling the geometric registration problem in a CNN framework and demonstrate superior registration accuracy and computational efficiency.
A Learning Strategy for Contrast-agnostic MRI Segmentation
3 code implementations • MIDL 2019 • Benjamin Billot, Douglas Greve, Koen van Leemput, Bruce Fischl, Juan Eugenio Iglesias, Adrian V. Dalca
These samples are produced using the generative model of the classical Bayesian segmentation framework, with randomly sampled parameters for appearance, deformation, noise, and bias field.
 Ranked #1 on
Brain Segmentation
on Brain MRI segmentation
Ranked #1 on
Brain Segmentation
on Brain MRI segmentation
FastSurfer -- A fast and accurate deep learning based neuroimaging pipeline
1 code implementation • 9 Oct 2019 • Leonie Henschel, Sailesh Conjeti, Santiago Estrada, Kersten Diers, Bruce Fischl, Martin Reuter
In this work we propose a fast and accurate deep learning based neuroimaging pipeline for the automated processing of structural human brain MRI scans, replicating FreeSurfer's anatomical segmentation including surface reconstruction and cortical parcellation.
Fast Infant MRI Skullstripping with Multiview 2D Convolutional Neural Networks
no code implementations • 27 Apr 2019 • Amod Jog, P. Ellen Grant, Joseph L. Jacobson, Andre van der Kouwe, Ernesta M. Meintjes, Bruce Fischl, Lilla Zöllei
The three probabilistic segmentations in the three views are linearly fused and thresholded to produce a final brain mask.
Unsupervised Deep Learning for Bayesian Brain MRI Segmentation
1 code implementation • 25 Apr 2019 • Adrian V. Dalca, Evan Yu, Polina Golland, Bruce Fischl, Mert R. Sabuncu, Juan Eugenio Iglesias
To develop a deep learning-based segmentation model for a new image dataset (e. g., of different contrast), one usually needs to create a new labeled training dataset, which can be prohibitively expensive, or rely on suboptimal ad hoc adaptation or augmentation approaches.
PSACNN: Pulse Sequence Adaptive Fast Whole Brain Segmentation
no code implementations • 17 Jan 2019 • Amod Jog, Andrew Hoopes, Douglas N. Greve, Koen van Leemput, Bruce Fischl
In this paper we propose a CNN-based segmentation algorithm that, in addition to being highly accurate and fast, is also resilient to variation in the input acquisition.
Pulse Sequence Resilient Fast Brain Segmentation
no code implementations • 30 Jul 2018 • Amod Jog, Bruce Fischl
We use the forward models to augment the training data with test data specific training examples.
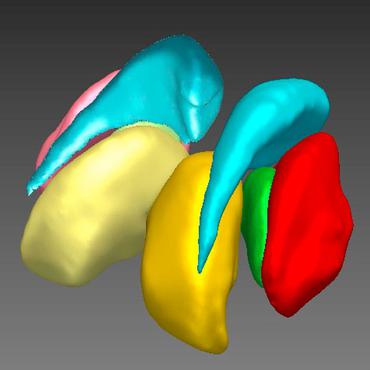
A probabilistic atlas of the human thalamic nuclei combining ex vivo MRI and histology
no code implementations • 22 Jun 2018 • Juan Eugenio Iglesias, Ricardo Insausti, Garikoitz Lerma-Usabiaga, Martina Bocchetta, Koen van Leemput, Douglas N. Greve, Andre van der Kouwe, Bruce Fischl, Cesar Caballero-Gaudes, Pedro M Paz-Alonso
In this study, we present a probabilistic atlas of the thalamic nuclei built using ex vivo brain MRI scans and histological data, as well as the application of the atlas to in vivo MRI segmentation.
Joint registration and synthesis using a probabilistic model for alignment of MRI and histological sections
no code implementations • 16 Jan 2018 • Juan Eugenio Iglesias, Marc Modat, Loic Peter, Allison Stevens, Roberto Annunziata, Tom Vercauteren, Ed Lein, Bruce Fischl, Sebastien Ourselin
Here, we overcome this limitation with a probabilistic method that simultaneously solves for registration and synthesis directly on the target images, without any training data.